
Ezymatic Triggered Self-Assembly and Gelation of ß-Sheet Peptides
Molecular
self-assembly has emerged as a powerful tool for the fabrication of
novel soft materials with wide ranging properties. In recent years,
considerable advances have been made in using self-assembling
oligopeptides as the building blocks for the preparation of hydrogels.
More recently the use of external stimuli such as light, pH, and ionic
strength to trigger their self-assembly has attracted considerable
attention. Enzymes is one other route to trigger the self-assembly of
peptides and proteins and to control the fabrication process of these
materials.
In the case of peptides, enzymes are usually used to convert a nongelling precursor into a self-assembling peptide. This common strategy consists of taking a well-known self-assembling peptide and modifying it with a side, or end-group that prevents its self-assembly, and consequently its gelation. The enzyme is then used to cleave the end/side group resulting in the self-assembly of the peptide and gelation of the sample. One other approach that has been used exploits the reverse hydrolytic properties of some enzymes. In this case, the enzyme is used to synthesize self-assembling peptides from a non self-assembling shorter precursor.
The
main focus of this project is to develop enzymatic trigger for the
production of hydrogels for biomedical applications exploiting the
reverse hydrolysis properties of thermolysine. Short
peptide usually 4 amino acid long that do not self-assemble in the
concentration range investigated are synthesised. Thermolysine is then
used to catalyse the synthesis of longer sequense mainly octa-peptides
that spontaneously self-assemble and subsequently result in the
gelation ofthe sample.
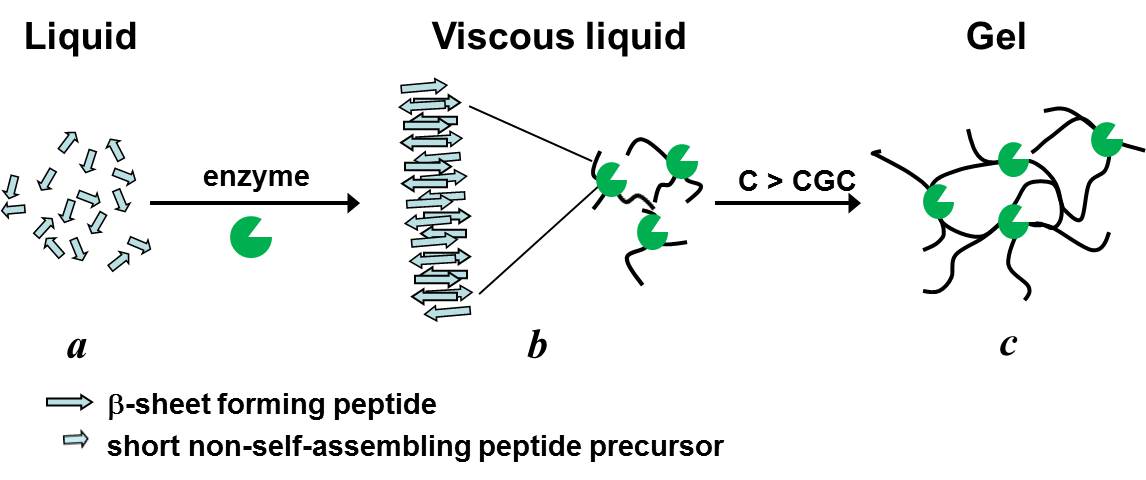
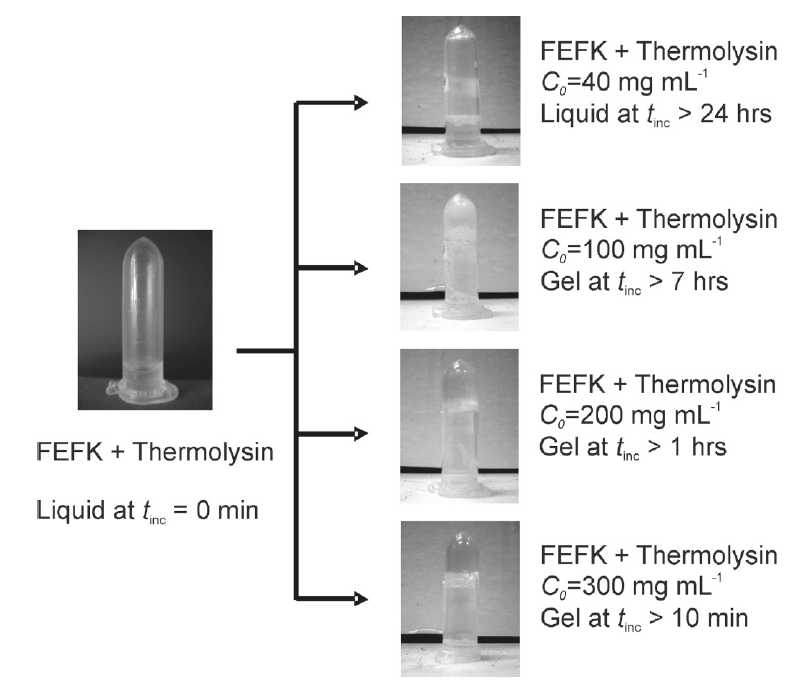
Left: Shematic representation of the enzymatic catalysis of self-assembling peptides from shorter precursor and their subsequent self-assembly and gelation. Right: Optical photographs of samples' macroscopic appearance (solution/gel) before and after the addition of thermolysin.
The properties, structural and mechanical, of hydrogels obtained through enzymatic catalysis are characterised using a variety of techniques including transmission electron microscopy (TEM), small angle X-ray scattering (WAXS), infrared spectroscopy (ATR-FTIR) and dynamic rheology. Using this novel approach hydrogel with signficant higher modulus can be obtained.
Left: Evolution of the storage, G', (closed symbols) and loss,G'', (open symbols)moduli of samples with initial tetrapeptide FEFK concentration of 40 mg mL-1 (circle), 100 mg mL-1 (diamond), and 200 mg mL-1 (square) after addition of the enxyme as a function of incubation time. Right: Sample storage moduli, G', after 2 h of incubation as function of the initial tetrapeptide concentration,C0.
These high mechanical properties derive from the heterogeneous structure of the networks formed through this appraoch. The self-assembling peptides are synthesized at the enzyme sites resulting in a local increase in concentration around the enzyme. These peptides self-assemble readily and form networks with higher density around the enzymes. As time goes on and more fibres are produced they extend throughout the sample. As a result of this process a heterogeneous networks are formed with denser “patches” being found around the enzymes.
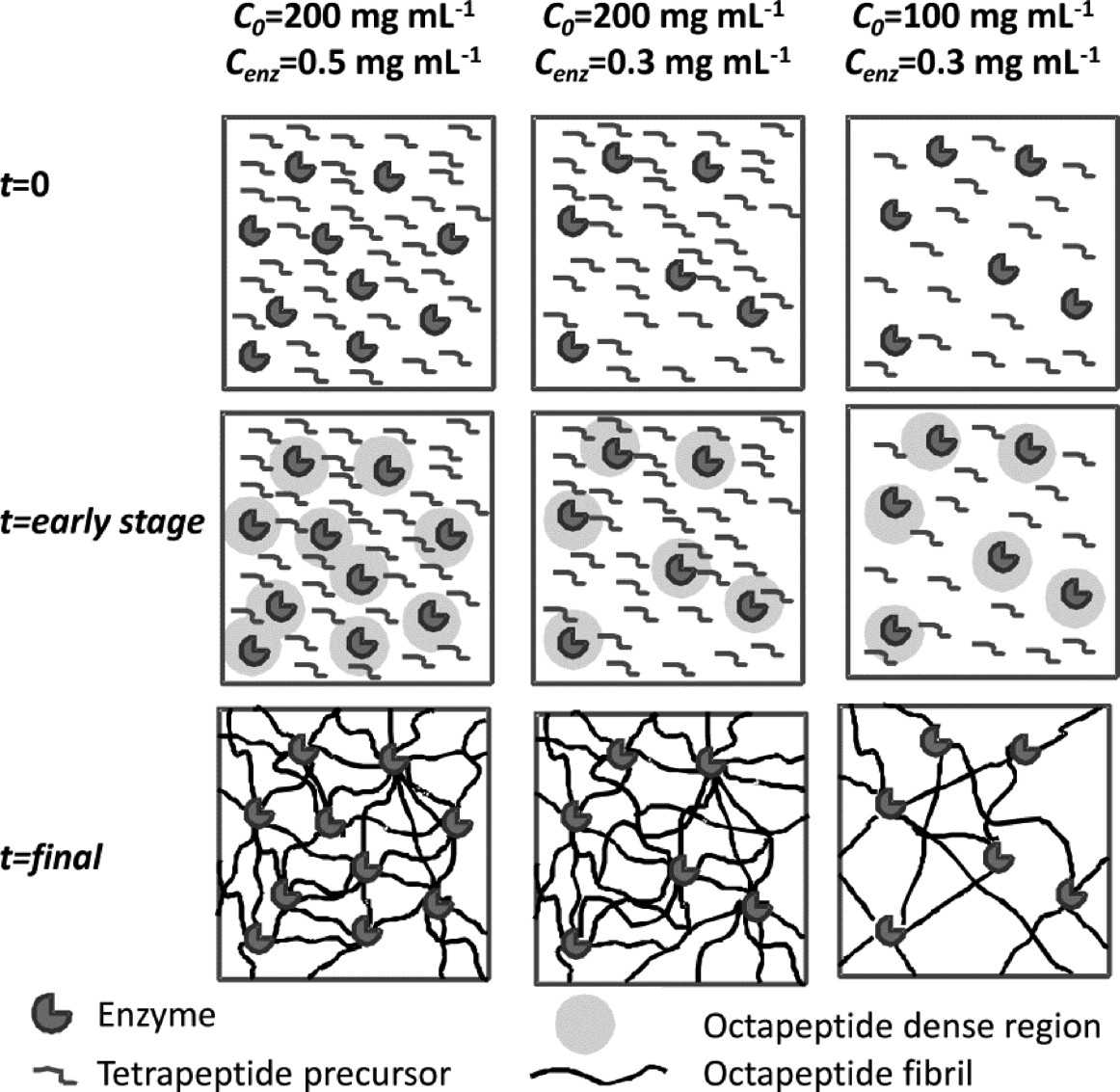
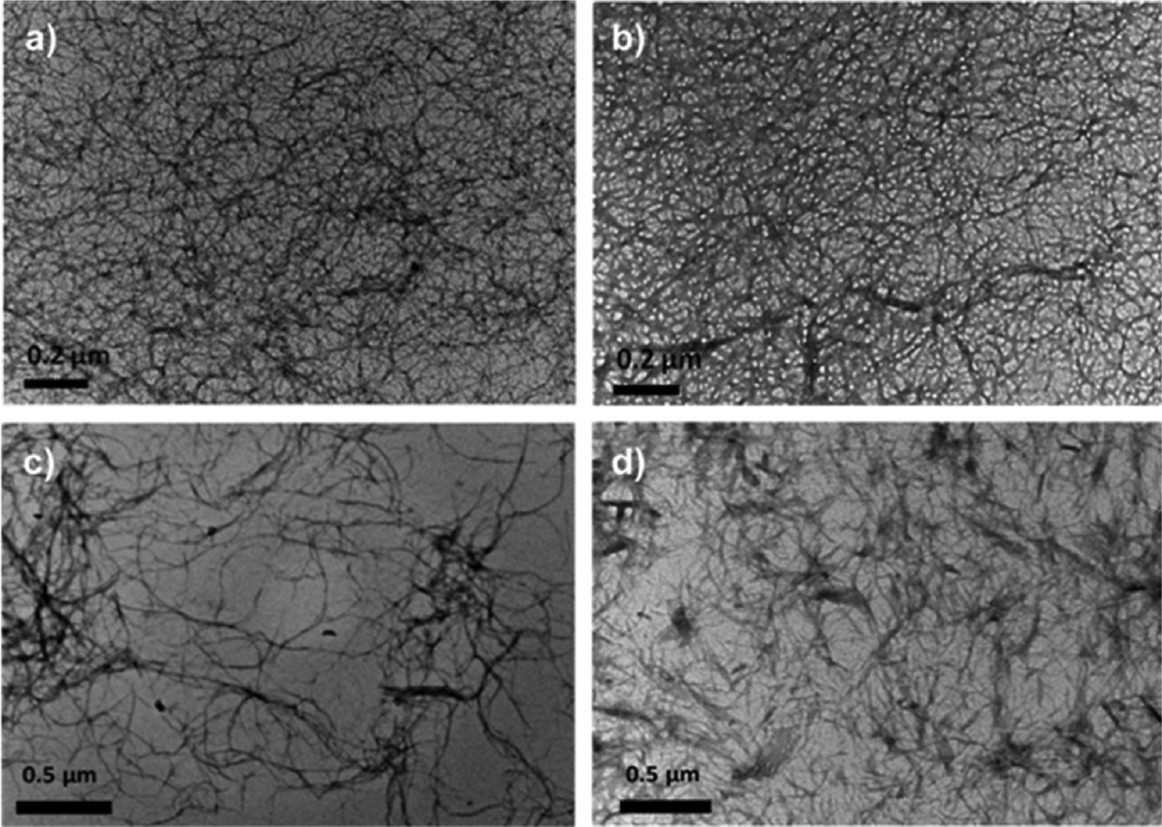
Left: Schematic representation of the fibrillar network of enzymatic triggered hydrogels. Right: TEM micrograph obtained for Cenz = 0.3 mg mL−1 samples: “non-diluted” after 20 min incubation (a) C0 = 100 and (b) 200 mg mL−1 and “diluted” after 1 day incubation (c) C0 = 100 and (d) 200 mg mL−1.
These denser network patches
have a
reinforcing role, as far as the hydrogel mechanical properties are
concerned, as they will have intrinsically higher modulus compared to
the rest of the gel. This type of reinforcing properties is akin to
composite materials to which fillers are added to reinforce them. This enzymatic gel preparation
method was shown to be suitable for the preparation of hydrogel for
cell culture.
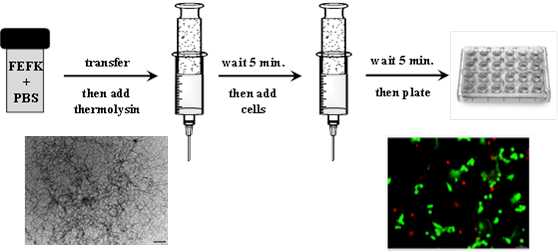
Use of enzymatically triggered peptide hydrogels for the 3D encapsulation and culture of cells
Publications:
________________________________________